13 August 2013
Coping with Jumping Junk
Genes determine and control traits by producing protein molecules. Some genes, however, do not appear to code for anything. The eukaryotic genome contains a lot of this “junk DNA”, which consist of DNA sequences that do not have a known functional role. As a matter of fact, approximately 98% of the human genome is made of junk DNA.
Where then, did all of this junk DNA come from? Much of junk DNA originates from DNA sequences called transposable elements (TE), which are also known as “jumping genes.” Unlike other DNA sequences, transposable elements jump around and insert themselves randomly in other parts of the genome. These insertions are often deleterious because they could alter gene expression in a way that induces genetic disease.
How do we suppress the deleterious effects of TEs? In a paper published on Nature Communications on August 12th, Prof. Saze of OIST's Plant Epigenetics Unit and his collaborators announced the discovery of a cellular mechanism in the flowering plant Arabidopsis that masks the effects of TEs. In particular, a protein they named IBM2 binds to TEs allowing genes to be expressed normally.
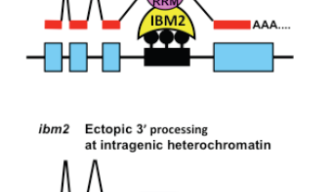
In a normal situation, genes are transcribed into messenger RNA (mRNA), which serves as a code for producing proteins. When a TE jumps into a gene, it alters the genetic code so that the mRNA no longer codes for the original protein. To counteract this, defense mechanisms that are comparable to the immune system come to work. Molecules are added on to the intruding TE sequence so that it becomes a heterochromatin, which is an inactive region of DNA. Simply turning the TE to a heterochromatin, however, is insufficient to produce a normal mRNA since the additional molecules on the TE block mRNA synthesis. Something else has to happen in order to produce mRNA beyond the heterochromatin.
This is where the IBM2 protein comes into play. IBM2 is a protein that binds to the inactivated TE. By doing so, it places a cover over the heterochromatin. This allows the transcription machinery to ignore the TE sequence and proceed as though the TE had not inserted itself. As a result, normal mRNAs will be transcribed despite having the heterochromatin which would normally interfere with mRNA transcription.
According to Prof. Saze, having mechanisms to overcome such obstacles allows for the evolution of complex gene controls in eukaryotes. This discovery deepens the understanding of how gene control mechanisms originated.
Specialties
Research Unit
Submit a press inquiry