30 June 2020
Breaking the silence: scientists investigate epigenetic impact across whole genome
All life depends on a genome, which acts as an instruction manual for building all the products essential for development and survival. But knowing which of these individual instructions – or genes – need to be read, and when, is key for a properly functioning organism: so how does life get this right?
Enter epigenetic regulation – the process by which cells control the expression, or readability, of genes. In multicellular organisms, epigenetics is the reason why every type of cell varies in shape and function, with each cell type following a different subset of instructions. Cells also use epigenetic regulation as an ‘immune system’, suppressing the activity of disruptive ‘jumping genes’ called transposons that can otherwise hop around the genome and threaten its integrity.
Despite its importance, scientists are still struggling to untangle the many pathways that cells use to precisely control the activity of genes. Now, researchers from the Okinawa Institute of Science and Technology Graduate University (OIST) have uncovered a clue to the mystery, by looking at how plant cells suppress transcription – the first stage of how genes manufacture their products. Their findings, recently published in Nature Communications, pinpoint previously unknown sections of DNA that are silenced by epigenetic regulation, many of which originate within transposons.
“This study provides a comprehensive view on how and where cells suppress transcription across the whole genome,” said Dr. Tu Le, first author and postdoctoral researcher in the OIST Plant Epigenetics Unit. “Importantly, we found that this silencing was vital for ensuring that genes involved in development and stress responses function properly.”
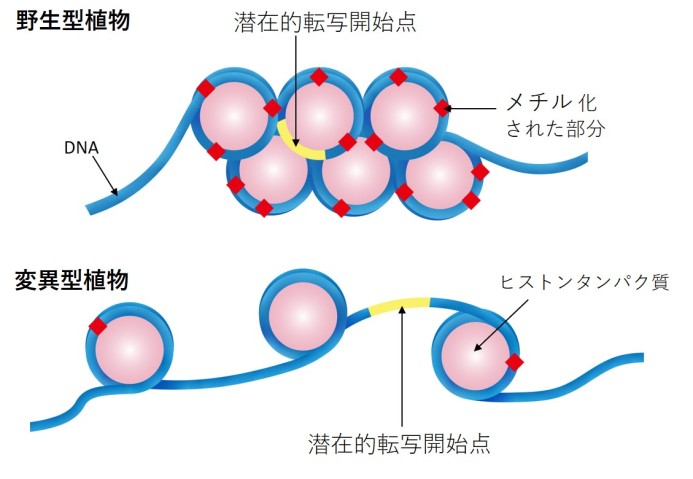
During transcription, cellular machinery copies a section of DNA into RNA. Usually, these RNA transcripts are then used to make proteins. Cells can boost or suppress transcription by adding chemical tags to DNA or to histone proteins that package the DNA, which tell the machinery which RNA transcripts – and ultimately proteins – to produce and in what quantity.
This level of precise control is vital for managing transposons. “Transposons are parasites of genomes, that promote their own expression at the expense of the organism,” said Professor Hidetoshi Saze, senior author of the study and leader of the Plant Epigenetics Unit. “When a transposon is active, its genetic sequence is used to manufacture a protein that can move the transposon to a different location in the genome, like cut-and paste or copy-and-paste computer functions.”
Transposons are usually silenced, as their activity can disable important genes. But sometimes, when under stress, plants re-activate transposons as they are a source of genetic variation, potentially generating beneficial mutations that allow the plant to adapt to the changing environment.
“Our lab ultimately aims to determine exactly how cells recognize and regulate transposons,” added Dr. Le. “This work is an important first step toward this goal.”
Unveiling hidden sites of transcription
In the study, the scientists used several mutant strains of a plant called Arabidopsis thaliana, with a different epigenetic pathway disabled in each strain.
The team then used a sequencing technique to detect specific DNA sequences that act as starting sites for the genome’s transcription machinery. They revealed thousands of these ‘transcription start sites’ (TSSs) that were only active in the epigenetic mutants.
“Many of these sites hadn’t been detected in previous studies, because they are completely silenced in wildtype plants. Our discovery of these hidden – or cryptic – TSSs provides a valuable source of data for future epigenetic research in plants,” said Prof. Saze.
The scientists identified one mutant strain of the plant that activated an especially high number of cryptic TSSs. The gene this mutant was missing encodes a key protein which maintains DNA methylation. When methyl groups are added to DNA, this epigenetic tag triggers a biochemical pathway that causes histones to pack the DNA more tightly. This physically stops transcription machinery from accessing the regions of the genome that contain the cryptic TSSs.
“The sheer number of cryptic TSSs activated when DNA methylation is lost shows that it is a powerful and prevalent method of silencing,” said Dr. Le.
From transposons to stress tolerance
Another key finding was the link between transposons and cryptic TSSs. The scientists found that up to 65% of cryptic TSSs originated within these ‘jumping genes’, which were longer and more heavily methylated that transposons without cryptic TSSs.
“This suggests that transposons with cryptic TSSs are younger, intact and still able to jump around the genome, which is why they are silenced,” explained Dr. Le.
Strikingly, the scientists noticed that when the cryptic TSSs were activated in the epigenetic mutants, this changed the activity of nearby genes involved in stress and development. The scientists don’t yet fully understand the mechanism behind this impact, but the implications are intriguing.
“There is previous research that shows that over time, as transposons degrade, plants can adapt the TSSs in transposons for their own use, to regulate the activity of nearby genes,” said Prof Saze. “The effect of the activated cryptic TSSs on stress and development genes suggests that in the future, plants could use these TSSs to adapt to changing conditions.”
In future research, the scientists hope to learn more about these cryptic TSSs and how they affect the activity of nearby genes. “This study might help us to better understand how plants respond to environmental changes such as global warming, drought and nutrient degradation in soil. It may then be possible to develop new crops which are resistant to these kinds of stress,” Prof. Saze said.
Specialties
Research Unit
Submit a press inquiry