03 July 2018
Mutation and 3D Modelling Reveal New Structure to Cell Division Process
Cell division is one of the most fundamental processes of biology. It’s what allows all living things to grow, multiply and form complex structures. But scientists still don’t understand many of the details behind the process.
Before a cell divides, it makes two copies of each of its chromosomes. These copies, called sister chromatids, align – and segregate -- to opposite ends of nucleus so that a full set will accompany the two daughter cells that will form. In a new study published in Proceedings for the National Academy of Sciences (PNAS), researchers at Okinawa Institute of Science and Technology Graduate University (OIST) are challenging a key aspect of how this process has been assumed to work.
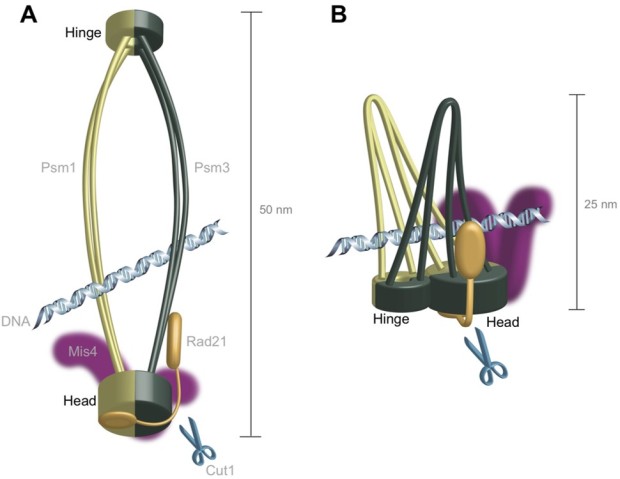
The study focuses on a protein complex called cohesin, which is tasked with holding the sister chromatids together until the cell begins to divide, and then letting go when the cell divides and the chromosomes must separate. Most researchers think that cohesin is shaped like a ring that holds chromatids together by encircling them. But for Drs. Xingya Xu and Norihiko Nakazawa at the OIST G0 Cell Unit headed by Professor Mitsuhiro Yanagida, some aspects of this model didn’t add up.
Cohesin releases its hold on chromatin with the help of an enzyme called Cut1. Chromatin is a mass of genetic material composed of DNA and proteins that condense to form chromosomes during eukaryotic cell division. When chromosomes are aligned, the Cut1 enzyme breaks open cohesin, allowing the two chromatids to segregate into the daughter cells. When the researchers mutated the gene encoding Cut1 in fission yeast cells, the cell division cycle broke down, as expected, because the mutated Cut1 couldn’t open the ring-shaped cohesin. However, in theory normal cell division could be restored by adding a second mutation that acts like an undo button for the first mutation, allowing the enzyme to break open cohesin once more, opening the ring.
That’s not what the team saw when they added this second mutation to their fission yeast cells. Instead, the cells that carried both these mutations were still not able to break down cohesin. Because the Cut1 enzyme needs cohesin’s assumed ring structure to function correctly, the scientists began to think that cohesin did not have a ring shape like researchers had long thought.
“We were surprised,” said Dr. Nakazawa, “because if cohesin was like a ring around chromatin, then the chromatids should have been cut loose by the second mutation.” The fact that it wasn’t, he adds, suggests that cohesin may have a different structure.
Previous studies in 2003 by Prof. Yanagida’s group had examined some aspects of cohesin’s structure at an atomic level using an extremely high-resolution microscope called an Atomic Force Microscope (AFM), but those results didn’t reveal all the fine details of the complex’s shape. To get a better picture, the OIST team had to look at the genetics.
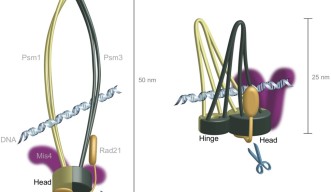
The scientists first sequenced the whole genome of cohesion in a fission yeast cell to identify where mutations occurred in areas where proteins interact with it, and how these mutations affect the structure of the cohesin complex. The mutations arose in areas of cohesin known as the “head” and “hinge”, which fasten the two main units of the protein complex together. They are thought to be on opposite sides of the ring structure, but further genomic analysis of cohesin itself suggested that they might be located next to one another, which would make cohesin’s ring configuration unlikely.
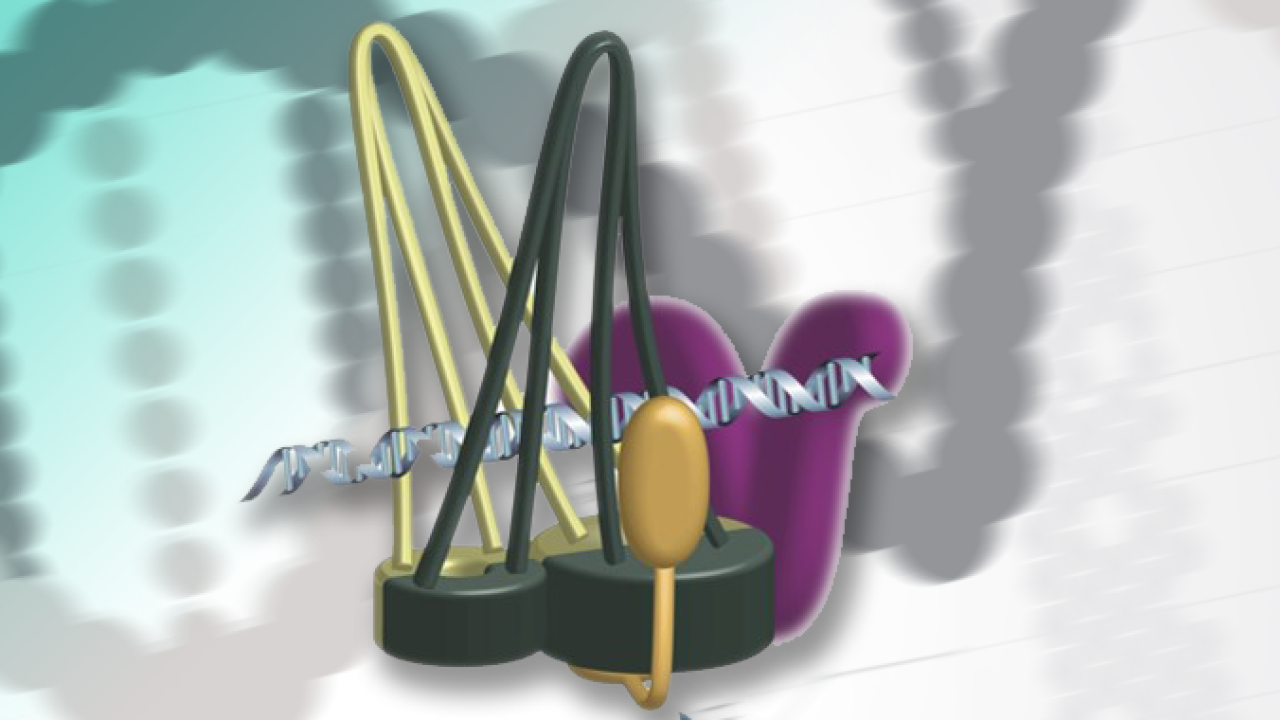
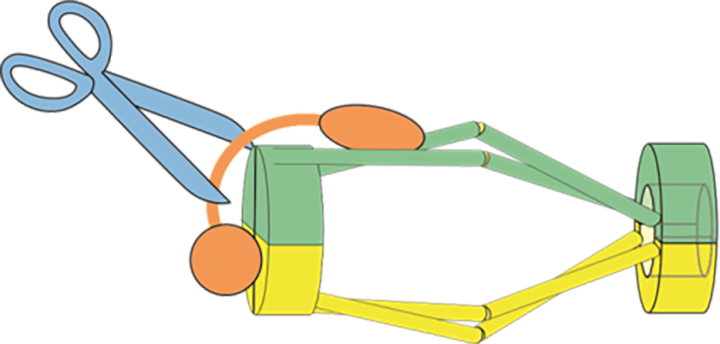
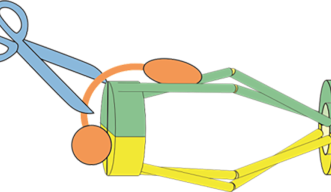
Prof. Yanagida, Dr. Xu and Dr. Nakazawa proposed a new model, dubbed hold and release, in which cohesin is more like a jaw that holds chromatids in place and then opens to allow chromatin to move, without the need for Cut1 to break open the complex.
Based on the sequences of cohesin’s head and hinge, the researchers constructed a 3D computer model of the protein complex. The model, created by Dr. Ryuta Kanai and Prof. Chikashi Toyoshima at the University of Tokyo, showed that the second mutation in the OIST fission yeast study couldn’t break down cohesin – supporting their observations in fission yeast and adding more evidence that the complex was not ring-shaped.
The new model fit well with AFM images of cohesin from the older study, which had suggested that the molecule was half the length of the expected ring structure and that cohesin was “tadpole shaped” – large at one end and tapering down to a thinner, opposite end.
Dr. Nakazawa and Dr. Xu still don’t have enough evidence to pin down cohesin’s shape. They also still don’t know how cohesin binds to chromatin to hold it in place and whether a single complex or a pair of them is needed to do so. As the team explore these possibilities, they may get closer to proving their new theory – and perhaps even glean a better understanding of diseases caused by abnormal cell division
Specialties
Submit a press inquiry