07 August 2013
Computational Models of Gene Regulation in Entire Fruit Fly Embryo
Although genomes are being sequenced with increasing frequency, it is not well understood how the genome specifies the mechanisms for forming a multicellular animal from a fertilised egg. Research Scientist Dr. Garth Ilsley, in Prof. Nicholas Luscombe’s Genomics and Regulatory Systems Unit at OIST, has been tackling this problem in collaboration with Assistant Professor Angela DePace at Harvard Medical School. He has succeeded in modelling early regulation of development in the fruit fly Drosophila using a statistical approach that accurately predicts gene expression in the whole embryo. This study was published in the open access journal, eLIFE, on August 6th.
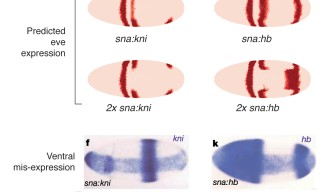
One of the genes essential for segmentation in the fly is called even skipped (eve). At an early stage of development, the eve gene can be detected as seven stripes around the embryo. Like all genes, its expression is repressed or activated by proteins called transcription factors. Since previous models could not explain the expression of the eve gene in the entire embryo of about 6,000 cells, it was assumed that some important factors had not been found and were not integrated into the calculation. Dr. Ilsley, on the other hand, constructed a simpler model that predicts expression in every cell using transcription factor concentration data from the Berkeley Drosophila Transcription Network Project. Dr. Ilsley’s approach enables calculations for the entire embryo for the first time. In this study, calculations were focused on two regions of regulatory DNA controlling stripes 2, 3 and 7. The expression of eve was predicted with considerable accuracy. The model also correctly predicts the effect on expression of the eve gene resulting from mutation or misexpression of its regulating transcription factors (Figure 1).
This model is expected to promote new biological discoveries by verifying hypotheses that could not be tested previously, or by comparing experimental results with the statistical model. Dr. Ilsley hopes to further develop the model at OIST using other organisms such as the chordate Ciona.
Submit a press inquiry