24 July 2012
Two OIST Units Team Up to Probe Life’s Beginnings
Whether a human, an armadillo, or a sea slug, each of us begins life in much the same way: as a microscopic, formless blob of stem cells. As we grow, an intricate choreography of genetic controls begins to differentiate our cells, turning some genes off and others up to full-blast, eventually yielding fully-formed animals. But the exact nature of that choreography—how each cell “decides” what job to perform, when to divide, and when to quietly die off—has remained somewhat of a black box, because discerning what is happening on the DNA strands in individual cells is extremely difficult.
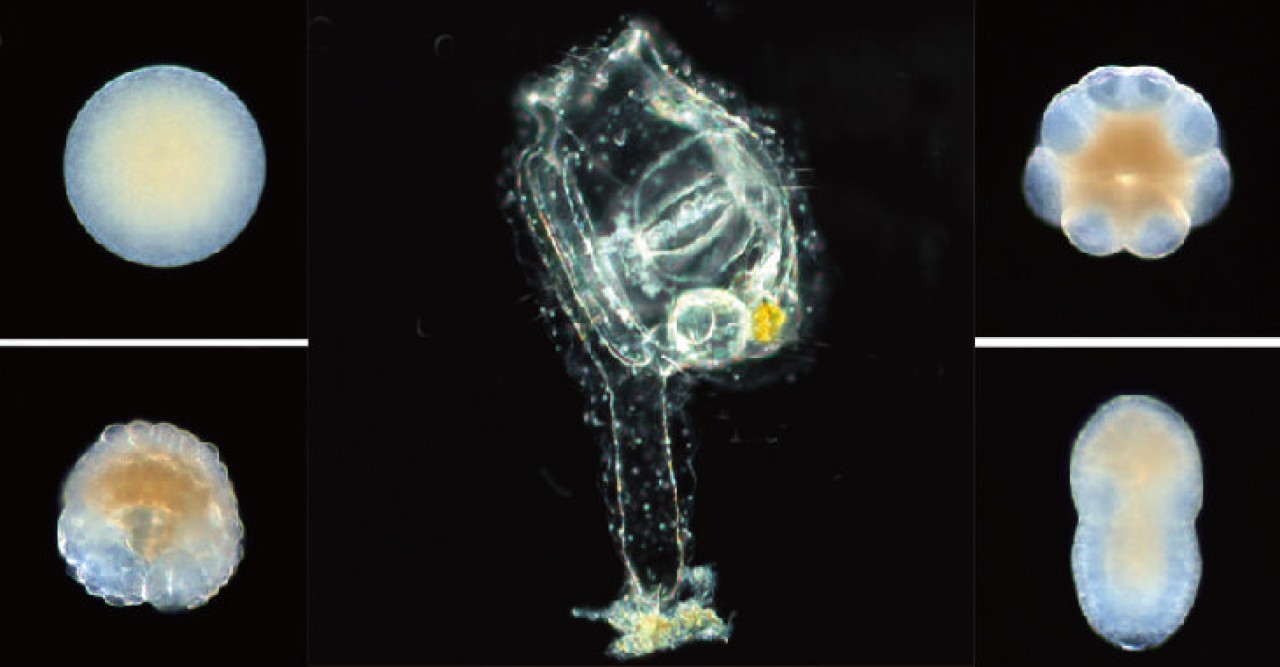
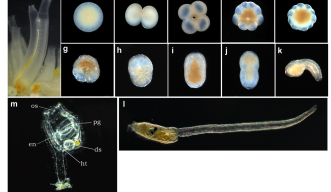
But Professors Nick Luscombe and Noriyuki Satoh are convinced that it can be done with the latest genetic sequencing and computational tools. Satoh, who heads the Marine Genomics Unit, will pick apart embryos of Ciona, a vase-shaped invertebrate sea animal, and measure gene activity in individual cells, while Takeshi Noda and Garth Ilsley in Luscombe’s Genomics and Regulatory Systems Unit will devise and run the computations that reveal what control mechanisms underlie the embryos’ march toward maturity.
The project will use embryos leading up to the 110-cell stage, when their development still closely parallels that of humans and other vertebrate animals, so the knowledge gained will have far-ranging implications. Ultimately, says Luscombe, “We would hope to be able to manipulate cells to make embryonic stem cells or certain tissues.”
Specialties
Research Units
Submit a press inquiry