02 April 2018
Biological Ballet: Development of a New Imaging Technique Reveals Complex Protein Movements in the Cell Membrane
What do ballet and cell biology have in common? Perhaps more than you might think.
Every cell in your body is enclosed by a cell membrane, a lipid bilayer that separates the cell’s contents from its surroundings. Residing within the cell membrane itself, molecules move around like ballet dancers on a stage.
In the words of Professor Akihiro Kusumi, leader of the Membrane Cooperativity Unit at the Okinawa Institute of Science and Technology Graduate University (OIST), “The proteins in the cell membrane undergo elegantly coordinated dances to relay messages between the cell and its environment.”
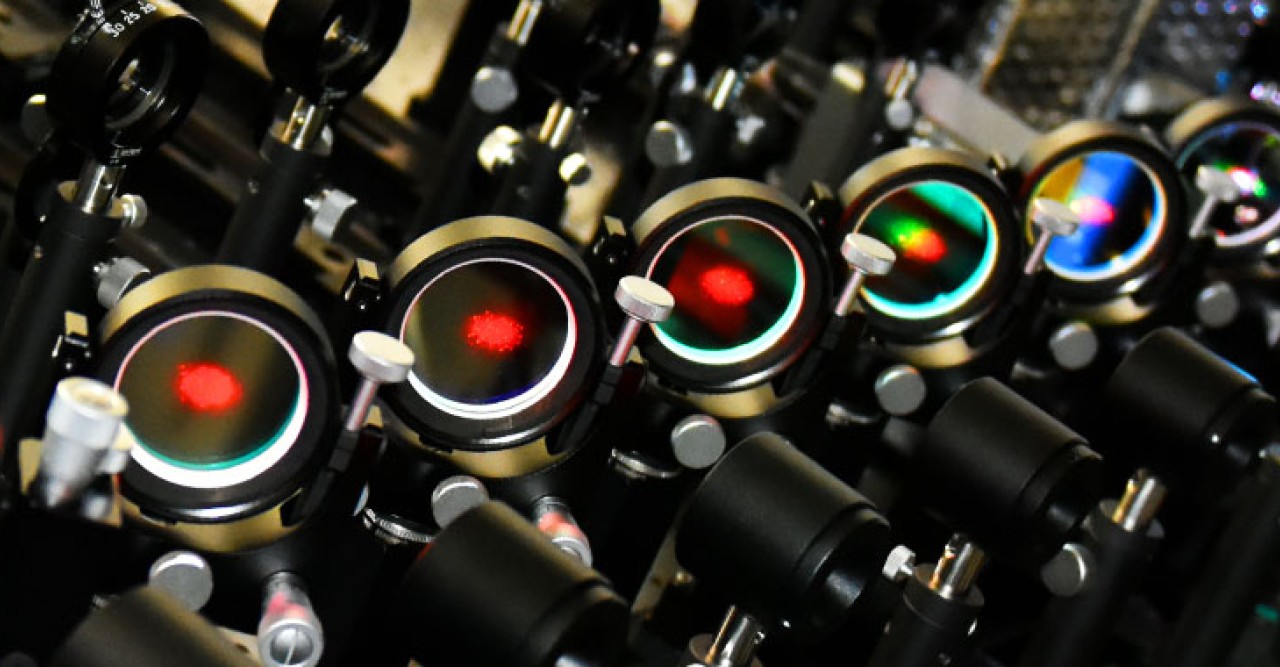
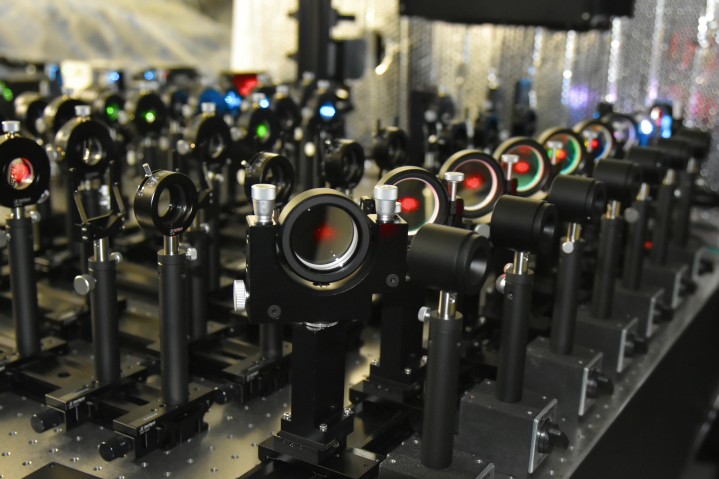
In order to understand how these proteins move within the membrane and how they interact with each other, Prof. Kusumi and other researchers developed an imaging method for live cells called live-cell single fluorescent-molecule imaging (SFMI). In SFMI, each protein ‘dancer’ in the membrane is individually tagged with fluorescent markers to make them visible under special home-built fluorescence microscopes.
However, SFMI can be problematic: — over time under the microscope, fluorescent markers lose their glow – a process known as ‘photobleaching’. Because of this, until now, cell biologists have been unable to observe individual molecules for longer than about ten seconds at a time. “It was like randomly taking many 10-second long clips, and trying to connect them in the correct order to produce a movie lasting for 5 minutes,” says Mr Taka-aki Tsunoyama, a researcher from the Membrane Cooperativity Unit at OIST.
In a paper recently published in Nature Chemical Biology, Tsunoyama, Kusumi, and their colleagues reported that they have come up with a twist on SFMI that suppresses photobleaching. Their method involves including a unique combination of chemicals and molecular oxygen in the specimen.
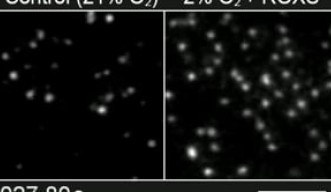
Previous methods used to prevent photobleaching were not very effective, and in addition, they were usually toxic to living cells, like totally removing molecular oxygen. However, the OIST researchers placed the cells in an environment with low oxygen concentrations, which mimics the real conditions inside a living organism, and added two mild chemicals called ‘trolox’ and ‘trolox quinone’. This combination was amazingly effective at reducing photobleaching without influencing the viability of the cells.
This new approach allows the researchers to observe individual molecules in living cells for up to 400 seconds. “Our method improves the observation time of fluorescent molecules by forty-fold,” says Prof. Kusumi.
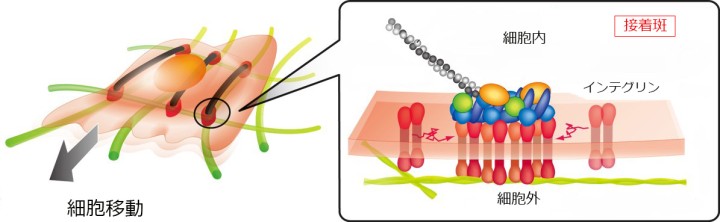
With the increased time window of observation, the researchers were able to study how molecules work in the cell far more directly and clearly. They studied specific regions of the membrane known as focal adhesions. “These are effectively the feet of the cell,” says Prof. Kusumi. The cell uses these ‘feet’ to get around, for instance, when cancer cells metastasize.
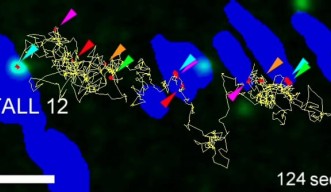
The researchers studied a group of proteins within the focal adhesion called integrins, which link the internal skeleton of the cell to the extracellular matrix. Previously, it was assumed that integrins are solidly fixed within the cellular feet. However, the extended observation time allowed the researchers to clearly see that integrin molecules repeatedly move and stop within the cellular feet, and even migrate between feet.
Like a rock climber searching for a new hold, an integrin molecule moves around to find a possible new attachment point. Upon finding one, it temporarily binds to it to determine whether the new hold is stable. Once it is deemed stable, the integrin grabs on tightly and pulls the cell forward.
Using their new technique, the researchers can record videos of entire acts of protein dances in the cell membrane, without interruption. “With our method, we can now follow the movement of each individual molecular dancer for sufficiently long periods of time to understand the cellular context” says Prof. Kusumi. Shedding light on the behavior of focal adhesion proteins could help researchers one day develop drugs that stop cancer cells from migrating through the body, he says.
Specialties
Research Unit
Submit a press inquiry