23 October 2018
Asian Catfish Genome Decoded
The striped catfish Pangasianodon hypophthalmus makes its home in the legendary Mekong River, the longest river in Southeast Asia and largest inland fishery in the world . Vietnam stands as the leading producer of the species, culturing an estimated 1.1 million tons of the fish in a single year. But unlike other commercial fish, such as the Atlantic cod or channel catfish, little genomic data exists to guide striped catfish aquaculture.
Now, scientists at the Okinawa Institute of Science and Technology Graduate University (OIST) have teamed with researchers at the Vietnam Academy of Science and Technology to decode the striped catfish’s entire genome. Their results, published October 5, 2018, in BMC Genomics, reveal new details about the species’ evolutionary lineage and genes that may be key to its healthy development.
“With this genomic data, it will be easy to check the diversification present in individual catfish populations,” said Eiichi Shoguchi, group leader of the Marine Genomics Unit at OIST and co-first author of the study. “In aquaculture, for example, an individual fish may be more resilient against disease than another. Now, you can check that individual’s genome and learn which gene is related to disease resistance.”
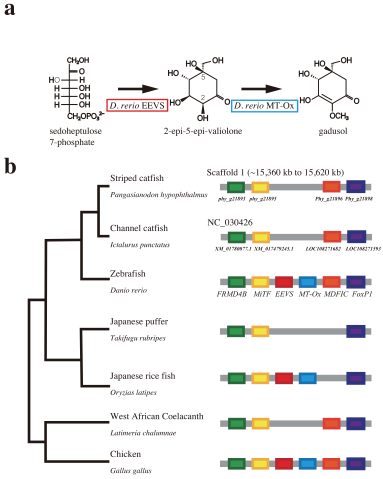
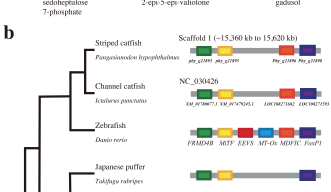
Using next generation sequencing techniques, the researchers gathered high-quality genomic data and assembled it into a complete genome. They compared the new genome to previously published genomic data from channel catfish and zebrafish -- a model organism closely related to catfish, evolutionarily. The scientists found distinct differences between the Asian catfish and its American cousin, and further differences between it and its relative the zebrafish.
In particular, they identified the loss of two genes in the striped catfish that remain intact in the zebrafish and allow the species to produce its own sunscreen. Catfish feed at the bottom of rivers, beyond the reach of UV-rays, and likely don’t require this protection. The striped catfish does have more insulin-like growth factor genes than zebrafish, which may be key to its healthy growth and development.
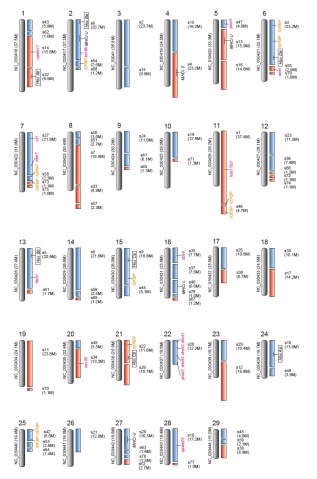
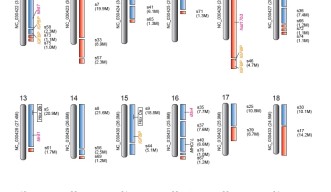
Using data from channel catfish, the scientists crafted a hypothetical map of striped catfish chromosomes -- the thread-like scaffolding where the fish’s genes are housed. The map points the way to future research comparing the species, tracing their lineage and investigating their gene function. Evolutionary biologists and aquaculturists both stand to learn from the striped catfish genome, now that it’d been decoded.
In the immediate future, aquaculturists can begin to seek out and select for optimal DNA fragments, known as “molecular markers,” into their own striped catfish chromosomes, said Oanh T. P. Kim, a lab head in the Institute of Genome Research at the Vietnam Academy of Science and Technology and co-first author of the study.
"The striped catfish genome information will enable us,” she said, “to develop these molecular markers and subsequently apply them in molecular marker-assisted selection for traits of economic importance in the catfish aquaculture."
Research Unit
Submit a press inquiry