Physics and Biology Unit
Principal Investigator: Jonathan Miller
Research Theme: Evolutionary, Comparative, and Biomedical Genomics
Abstract
Darwin postulated that evolution proceeds by the action of natural selection on neutral variation. Comparative genomics aims to disentangle the effects of natural selection on genome sequences from those of neutral variation. Although this program has achieved enormous success in medicine and fundamental biology over the last fifty years, estimates of functional genome sequence in human jumped from just over 2% to around 6% on the comparison of the whole genome sequences of human and mouse in 2002. What about the other 94% ?
Our Unit has accumulated evidence over recent years that an incomplete understanding of neutral variation is hindering further advances. In particular, although it is generally recognized that treating neutral variation as a process dominated by uncorrelated base substitutions is unrealistic, no practical alternatives have been proposed that are consistent with sequence data.
To develop such alternatives, our Unit takes a data-based approach to the problem through physics-style phenomenology and data analysis. Our calculations suggest that strong sequence conservation among diverse species can arise from sources other than selection. One possible source is the structure of neutral variation, which is likely to be more complex than generally appreciated.
1. Staff
- Dr. Kun Gao, Researcher
- Dr. Sathish Venkatesan, Researcher
- Dr. Maxim Koroteev, Researcher
- Dr. Eddy Taillefer, Researcher
- Midori Tanahara, Research Administrator
2. Collaborations
N/A
3. Activities and Findings
3.1 Comparative Genomics and Neutral Variation
Darwin told us that evolution is (adaptive) selection acting on (random) variation. Darwin didn't know about DNA. But we now know that DNA sequence variation is a major source of the variation that Darwin described. Following Darwin, “neutral" sequence variation is the variation that would be observed in the absence of selective pressure on the sequence.
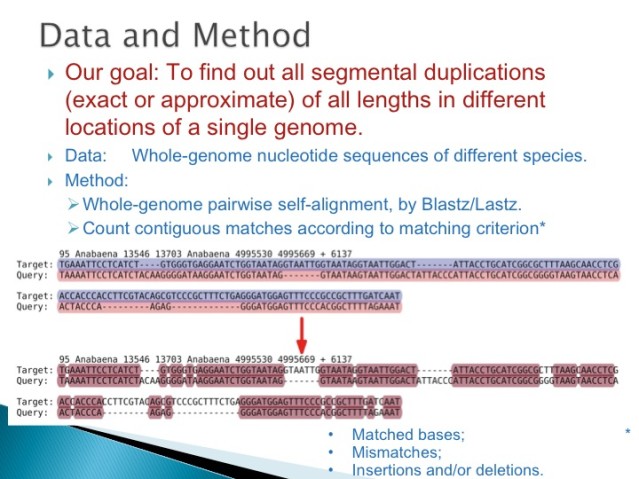
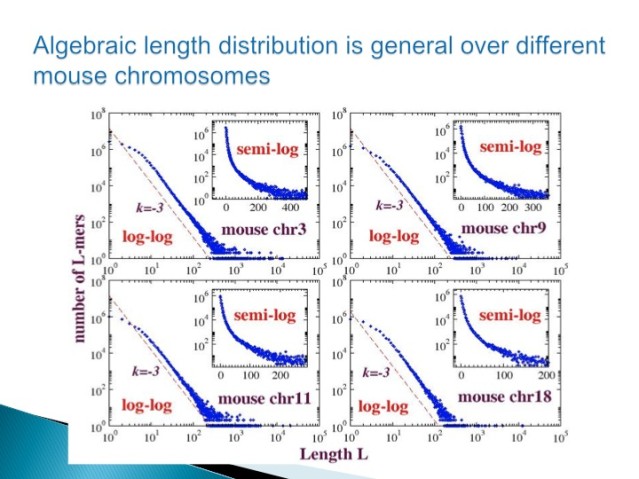
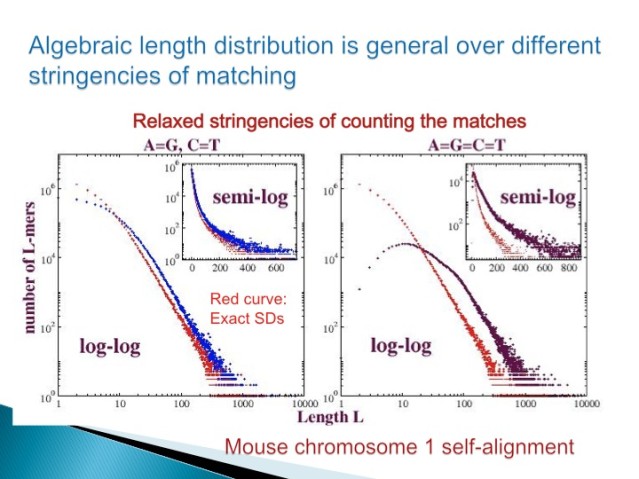
3.2 Duplication is One Form of Neutral Variation
Figure 3.
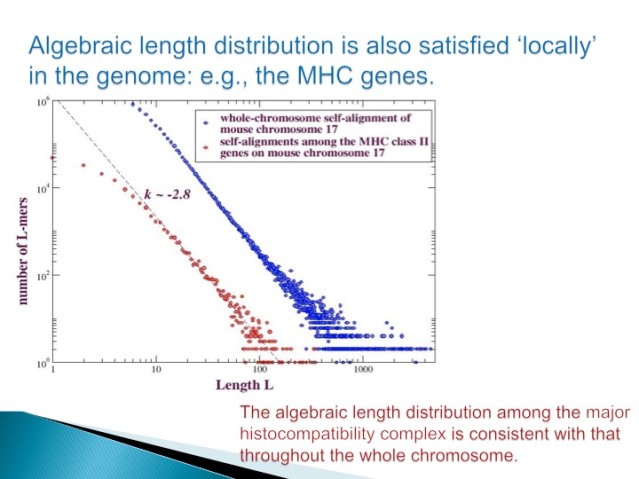
Even more dramatically, we don't have to look at whole chromosomes to see this distribution - a large gene family such as the major histocompatibility genes - is sufficient, as illustrated in Figure 4.
Figure 4.
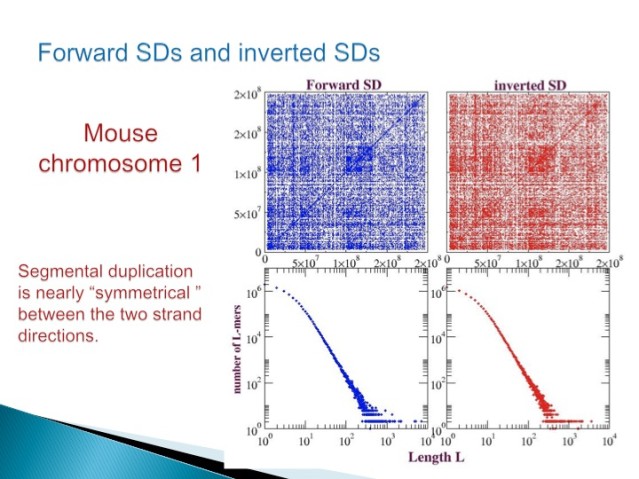
Finally, both forward and inverted duplications conform to the algebraic distribution (Figure 5).
Figure 5.
4. Publications
4.1 Journals
Submitted (2010):
1. Koroteev, M. & Miller, J. Scale-free Duplication Dynamics: A Model for Ultraduplication. (Accepted, Physical Review E, August 2011).
2. Gao, K & Miller, J. Algebraic Distribution of Segmental Duplication Lengths in Whole-Genome Sequence Self-Alignments. (Accepted, PLoS ONE, March 2011).
4.2 Books and other one-time publications
N/A
4.3 Oral and Poster Presentations
- Gao, K. & Miller, J. Algebraic Distribution of Segmental Duplication Lengths in Whole-Genome Sequence Self-Alignments, Computational Biology, held by Cold Spring Harbor Asia, Suzhou China, Sep 27-Oct 1, 2010
- Taillefer, E. & Miller, J. Algebraic length distribution of sequence duplications in whole genomes, Computational Biology, held by Cold Spring Harbor Asia, Suzhou China, September 27-Oct 1, 2010
- Venkatesan, S. & Miller, J. Spatial correlations among the third bases of codons for perfectly-conserved amino acid coding sequences, Computational Biology, held by Cold Spring Harbor Asia (International), Suzhou China, Sep 27-Oct 1, 2010
- Koroteev, M. & Miller, J. A model for ultraduplication, Computational Biology, held by Cold Spring Harbor Asia, Suzhou China, Sep 30, 2010
- Miller, J. Intensive and exhaustive genome sequence comparison: lessons for biology and challenges for computation, IPAB Workshop, organized by NPO Initiative for Parallel Bioinformatics. "Seeds and Needs for Large Scale Computing 2010 -Next Generation Sequencer : Uniting IT and Biotechnology", Naha, Okinawa, Japan, Oct 1, 2010
- Miller, J. Algebraic sequence correlation arises from Recombination, Kavli Institute for Theoretical Physics, University of California, Santa Barbara, USA, Feb 16, 2011
5. Intellectual Property Rights and Other Specific Achievements
Nothing to report.6. Meetings and Events
6.1 Seminar
- Title: "Life without water: molecular mechanism to stand complete desiccation in the Sleeping Chironomid, Polypedilum vanderplanki"
- Date: June 16, 2010
- Venue: Lab 1, OIST
- Speakers: Takashi Okuda (National Institute of Agrobiological Sciences Tsukuba)
- Co-organizers: Noriyuki Satoh (OIST) and Alexander Mikheyev (OIST)
6.2 OIST Internal Seminar
- Title: "Distribution of segmental duplication lengths in whole genomes"
- Date: July 9, 2010
- Venue: Lab 1, OIST
- Speaker: Kun Gao (OIST)
6.3 International Workshop
- Title: "Quantitative Evolutionary and Comparative Genomics (QECG) 2010"
- Date: May 24 - June 4, 2010
- Venue: Seaside House, OIST
- Co-organizers: Holger Jenke-Kodama (OIST), Alexander Mikheyev (OIST) and Byrappa Venkatesh (IMCB, Singapore)
7. Others
N/A