Open Biology Unit
Principal Investigator: Hiroaki Kitano
Research Theme: Computational systems biology and software platform development for drug discovery and medicine
Abstract
Our unit aims to develop a novel software platform for systems biology and systems drug discovery that may fundamentally transform the way we study these fields.
Modern biology and medicine are fundamentally data- and evidence-driven, and researchers are fighting against the vastness and complexity of the data and the scattered nature of our knowledge. In order to obtain in-depth insights that may lead to new biological discoveries and medical implications, one must be able to properly access this data and knowledge and then integrate, analyse, and link them to practical solutions. This requires a new approach in science in the sense that it has to be open-ended, evolvable, community-based, and computationally supported.
Our unit is developing a methodology and software platform for open biology that entails the above features. This includes teh Payao community-based pathway curation system, PhysioDesigner physiological modeling tool, and a series of software packages that will be interoperable to The Garuda Platform. The Garuda Platform is a global alliance that we initiated to create a consistent, integrated user experience - a one-stop service style software platform. Such a platform will enable researchers to explore system-level characteristics of biological systems more effectively than before. At the same time, we are exploring a novel drug discovery approach based on computational systems biology, theory of biological robustness, and a concept of long-tail drugs.
1. Staff
- Dr. Yoshiyuki Asai, Group Leader
- Dr. Kun-Yi Hsin, Researcher
- Yuuki Tamashiro, Programmer
- Midori Tanahara, Research Administrator
2. Collaborations
- Theme: Software platform for systems biology
- Type of collaboration: Research Collaboration (no financial transaction)
- Researchers:
- Dr. Samik Ghosh, Systems Biology Institute
- Ms. Yukiko Matsuoka, System Biology Institute
- Theme: Integration of Manchester text mining system to Payao
- Type of collaboration: Research Collaboration (no financial transaction)
- Researchers:
- Prof. Sophia Ananiadou, Manchester University
- Theme: Open platform for multi-level modeling
- Type of collaboration: Research Collaboration (no financial transaction)
- Researchers:
- Professor Taishin Nomura, Osaka Univerisity
- Professor Hideki Oka, Osaka University
3. Activities and Findings
3.1 Development of a social curation platform enabling a community to work on pathway models.
We have been developing a web-based tool called Payao (http://celldesigner.org/payao) to enable a community to work on a same pathway mode together. Payao reads models in SBML (Systems Biology Markup Language; http://sbml.org) and displays them in SBGN (Systems Biology Graphical Notation; http://www.sbgn.org/) formats, fully complying with the community standards. Over the platform, users can insert tags to a specific part of the model (such as Species, Reactions and specified area), exchange comments, record the discussions and eventually update the models. It promotes knowledge integration and collaboration among different expertise groups by which models can be therefore comprehensively curated and optimized. The information given by users is stored in a database on server allowing data retrieval for further curation. Payao is also capable of handling the models of CellDesigner (http://celldesigner.org/), a structured diagram editor for drawing gene-regulatory and biochemical networks, parsing SBML files sent from client to create CellDesigner models. The curation data on Payao can accordingly be reintegrated into the original model via CellDesigner.
3.2 Toward open and united platform for systems biology and integrated life science
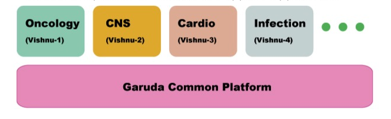
Numbers of tools and resources are being developed to foster systems biology and healthcare research. However, most tools are developed independent to each other. In most cases, there is no coordination with other groups. However no sufficient work has been initiated for interoperability and efficient data sharing. To resolve this problem, Garuda initiative was launched with the inaugural workshop took place in OIST in February 2010. After having two successive workshops in Manchester and Edinburgh, we had the fourth workshop again in February 2011 at OIST. Until now we have been discussing on the software architecture of the Garuda core, services that can be potential alliance member, and application APIs that should be provided by each service. We have defined primary specifications for the core architecture and APIs, and we have already started to develop prototypes according to the specifications.
Figure 1: Conceptual diagram of Garuda common platform
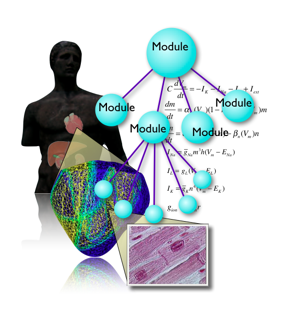
3.3 Development of tools for multilevel mathematical modeling for physiological functions
Accumulation of knowledge of physiology has opened a new scientific field, integrated life-science in which inter-level principles as well as intra-level disciplines are explored. Roles played by multi-scale and multi-level mathematical modeling of physiological functions are becoming more and more important. The framework for supporting to build such mathematical models of physiological functions, and for archiving and sharing models are inevitable for further development. We are developing a system called PhysioDesigner inheriting a previously developed platform insilicoIDE at physiome.jp, besides other pioneering efforts to promote physiome and systems biology, such as SBML and CellML. On our platform, physiological functions are considered as an aggregate of “modules” which are easily viewed and edited on PhysioDesigner. Based on this modularity, physiological functions are structuralized and modeled. The model is described in insilicoML (ISML), an XML based language, which we have defined already to describe the modular and hierarchical structure of the models. Using our platform users can integrate not only mathematical expressions but also experimentally obtained time-series data and morphological data. We are also extending a simulator which supports ISML and SBML. The most prominent feature of the simulator is its function supporting parallel computing. Users can concentrate on logics of models without bothering numerical calculation algorithms including parallelization.
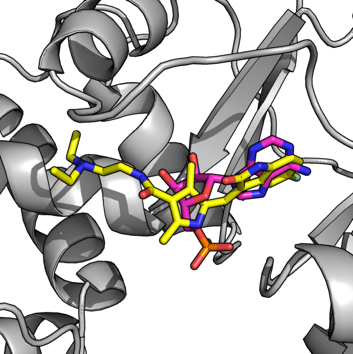
3.4 Application of systems biology and structural biology in drug discovery
Figure 3: Virtual docking for protein-ligand interaction study.
4. Publications
4.1 Journals
- Caron E., Ghosh S., Matsuoka Y., Ashton-Beaucage D., Therrien M., Lemieux S., Perreault C., Roux PP. & Kitano H. A comprehensive map of the mTOR signaling network. Mol Syst Biol 6, Article no. 453, doi:msb2010108 [pii] 10.1038/msb.2010.108 (2010).
- Ghosh S., Matsuoka Y. & Kitano H. Connecting the dots: role of standardization and technology sharing in biological simulation. Drug Discov Today 15, 1024-1031, doi:S1359-6446(10)00336-3 [pii] 10.1016/j.drudis.2010.10.001 (2010).
- Kaizu K., Ghosh S., Matsuoka Y., Moriya H., Shimizu-Yoshida Y. & Kitano H. A comprehensive molecular interaction map of the budding yeast cell cycle. Mol Syst Biol 6, 415, doi:msb201073 [pii] 10.1038/msb.2010.73 (2010).
- Kaizu K., Moriya H. & Kitano H. Fragilities caused by dosage imbalance in regulation of the budding yeast cell cycle. PLoS Genet 6, e1000919, doi:10.1371/journal.pgen.1000919 (2010).
- Kemper B., Matsuzaki T., Matsuoka Y., Tsuruoka Y., Kitano H., Ananiadou S. & Tsujii J. PathText: a text mining integrator for biological pathway visualizations. Bioinformatics 26, i374-381, doi:btq221 [pii] 10.1093/bioinformatics/btq221 (2010).
- Kitano H. Violations of robustness trade-offs. Mol Syst Biol 6, Article no. 384, doi:msb201040 [pii] 10.1038/msb.2010.40 (2010).
- Kitano H. Grand challenges in systems physiology. Frontiers in Physiology 1 (2010).
- Matsuoka Y., Ghosh S., Kikuchi N. & Kitano H. Payao: a community platform for SBML pathway model curation. Bioinformatics 26, 1381-1383, doi:btq143 [pii] 10.1093/bioinformatics/btq143 (2010).
- Shiraishi T., Matsuyama S. & Kitano H. Large-scale analysis of network bistability for human cancers. PLoS Comput Biol 6, e1000851, doi:10.1371/journal.pcbi.1000851 (2010).
- Kitano H., Matsuoka Y. & Ghosh S. Computational infrastructure for systems biology. Instrument and Control Engineers 49, 507-512 (2010).
4.2 Books and other one-time publications
None4.3 Oral and Poster Presentations
- Asai Y. Demonstration of PhysioDesigner as a partner of Garuda alliance, Garuda Four Workshop, Seaside House, OIST, Okinawa, Feb 23, 2011
- Kitano H. Garuda Alliance Perspective, Garuda Three Workshop, Informatics Forum building at the University of Edinburgh, Oct 9, 2010
- Kitano H. Garuda Alliance: Current status, Garuda-4 Workshop, OIST, Okinawa, Feb 23, 2011
- Kitano H. Hitting Robustness, 16th World Congress on Basic and Clinical Pharmacology (WorldPharma 2010), Bella Center, Copenhagen, Denmark, Jul 19, 2010
- Asai Y. Method and its application for physiome programing, Fifth Dynamic Brain Platform Committee, Chateraise Gateaux Kingdom SAPPORO, Sapporo, Dec 17, 2010
- Asai Y. Possible use case: CellDesigner and insilicoIDE (PhysioDesigner) Communication for Multilevel Modeling., Garuda Three Workshop, Informatics Forum, Edinburgh, UK., Oct 9, 2010
- Kitano H. Rationale of Garuda and introduction to Garuda-2 worksho, Garuda Two Workshop, Manchester Interdisciplinary Biocentre, UK, 07.05, 2010
- Kitano H. SBI software platform for systems biology, Sage Commons Congress, JW Marriott Union Square, San Francisco, USA, Apr 23, 2010
- Asai Y. and Villa A.E.P. Spatio-temporal filtering of the distributed temporal information in spike train through a diverging/converging neural network. , The 9th Intenational Neural Coding Workshop, Limassol, Cyprus. , Oct 30, 2010
- Kitano H. Standard formation in systems biology: past and future, SBML 10th Anniversary Symposium, Informatics Forum building at the University of Edinburgh, UK, Oct 9, 2010
- Kitano H. Systems Biology and Drug Discovery, Open Source Drug Discovery Connect to Decode Workshop, Teen Murti Bhawan, New Delhi, India, Apr 9, 2010
- Kitano H. Systems Biology and Drug Discovery, Talk at University of Luxembourg, University of Luxembourg, Luxembourg, May 3, 2010
- Kitano H. Systems Biology and Drug Discovery, National University of Singapore, Biochemistry Seminar, National University of Singapore, Singapore, Nov 8, 2010
- Kitano H. Systems Drug Design, Systems Biology at the Interface of Metabolism and Signal Transduction, VU University Amsterdam, The Netherlands, Dec 17, 2010
- Kitano H. Systems drug discovery and software platform for healthcare research and services, InCoB (International Conference on Bioinformatics), Waseda University, Tokyo, Sep 27, 2010
- Kitano H. Systems Medicine and Systems Drug Design as the Grand Challenge in 21st Century, BioComplexity Symposium: Systems Biology Approach to Systems Drug Discovery and Systems Medicine, NUHS Tower Block, Singapore, Feb 14, 2011
- Kitano H. Systems Robustness and Systems Failures in Biology, Medicine, and Engineering, A*STAR Scientific Conference 2010, A*STAR, Singapore, Nov 10, 2010
- Kitano H. Systems Biology and its application to drug design, 33rd Workshop on biological instrumentation), Toyo Corporation, Tokyo, Feb 7, 2011
- Kitano H. Global Strategy for Systems Biology Research, JST Seminar: Science and technology policy seminar, Center for Research and Development Strategy, May 14, 2010
- Kitano H. Systems Toxicology, 37th Annual Meeting of the Japanese Society of Toxicology, Okinawa Convention Center, Okinawa, Jun 16, 2010
- Ghosh S., Matsuoka Y. & Kitano H. An Open-flow Paradigm for Curation and Enrichment of Biological Pathways, ICSB 2010, Edinburgh International Conference Centre, UK, Oct 12, 2010
- Kemper B., Matsuzaki T., Matsuoka Y., Tsuruoka Y., Kitano H., Ananiadou S. & Tsujii J. PathText: Integrating Text Mining Resources with Biological Pathway Visualizations ICSB 2010, Edinburgh International Conference Centre, UK, Oct 12, 2010
- Kitano H. Systems Drug Design, Kanagawa Academy of Science and Technology(KAST) Course Systems Biology 2010, Kanagawa Academy of Science and Technology(KAST), Kanagawa, Nov 18, 2010
- Asai Y. Platform for Multi-level Modeling and Simulation, Brain-IS Workshop: : Integration of Brain Science and Engineering), Kyusyu Institute of Technology, Fukuoka, February 10, 2011
- Kitano H. The role of OIST in Okinawa Bio-industry cluster formation, Okinawa New Industry Formation Investment Program, Todofuken-Kaikan, Tokyo, March 2, 2011
5. Intellectual Property Rights and Other Specific Achievements
- JP2010-130513 (PCT/JP2011/058229)
- JP2010-224308 (PCT/JP2011/058323)
6. Meetings and Events
6.1 International Joint Conference of Artificial Intelligence (IJCAI) Trustee Meeting
- Dates: August 7-8, 2010
- Venue: Seaside House, OIST
- Sponsor: IJCAI (International Joint Conference on Artificial Intelligence)
- Committee:
- Craig Knoblock (University of Southern California)
- Toby Walsh (University of New South Wales)
- Sebastian Thrun (Stanford University)
- Francesca Rossi (University of Padova)
- Ramon López de Mántaras (Artificial Intelligence Research Institute/Spanish Scientific Research Council)
- Craig Boutilier (University of Toronto)
- Fausto Giunchiglia (University of Trento)
6.2 Garuda Four (Garuda Workshop Series)
- Dates: February 23-26, 2011
- Venue: Seaside House, OIST
- Sponsor: OIST, Systems Biology Institute, University of Tokyo, Manchester University
- Speakers:
- Huaiyu Mi (University of Southern California)
- Taishin Nomura (Osaka University)
- Natalia Polouliakh (Sony CSL)
- Akira Funahashi (Keio University)
- Venkata Satagopam (EMBL)
- Yukiko Matsuoka (JST ERATO)
- Richard Adams (University of Edinburgh)
- Reinhard Schneider (EMBL)