20 September 2022
Tracking a network of 100,000 mutants
Highlights:
- Mutations are responsible for most of the diversity of life we see around the world today, but many aspects of how mutations affect an organism’s fitness remain poorly understood.
- Now, researchers have experimentally shown, for the first time, a concept about mutations and evolution that has previously only been theoretically predicated.
- Called a neutral network, it’s thought to be vital for increasing genetic diversity in a population.
- A neutral network is a whole series of base sequences, where each new sequence, differing by just one base, has roughly the same fitness as the one before and after.
- The researchers emphasized that this will help scientists test many hypotheses, especially around how RNA evolved and continues to evolve.
Press release:
What fundamentally sets a human being apart from every other living creature comes down to differences in DNA sequences—a set of genetically-inherited molecules found in every cell of every organism. These differences have accumulated over millions of years, mainly via random mutations—basically errors in how the DNA was copied. Most of these mutations negatively impact the organism and will likely result in it dying before it has a chance to reproduce. However, some will positively, or neutrally, impact the organism and spread through the population. These mistakes in DNA sequences have resulted in the diversity of life we see around the world today. But many aspects of how these mutations can increase fitness remain poorly understood.
“We’ve managed to experimentally show, for the first time, a concept about mutation and evolution that has previously only been theoretically predicated,” said Prof. Yohei Yokobayashi, who leads to Nucleic Acid Chemistry and Engineering Unit at the Okinawa Institute of Science and Technology Graduate University (OIST). “It’s called a neutral network and it’s thought to be vital for increasing diversity in a population.” This research was published in Nature Communications.
Genes, made up of DNA base pairs, contain the instructions needed to create proteins, and lead to the proper care and maintenance of a cell. For the instructions to be carried out, the DNA must first be transcribed into RNA. Thus, RNA is like a reflection of DNA.
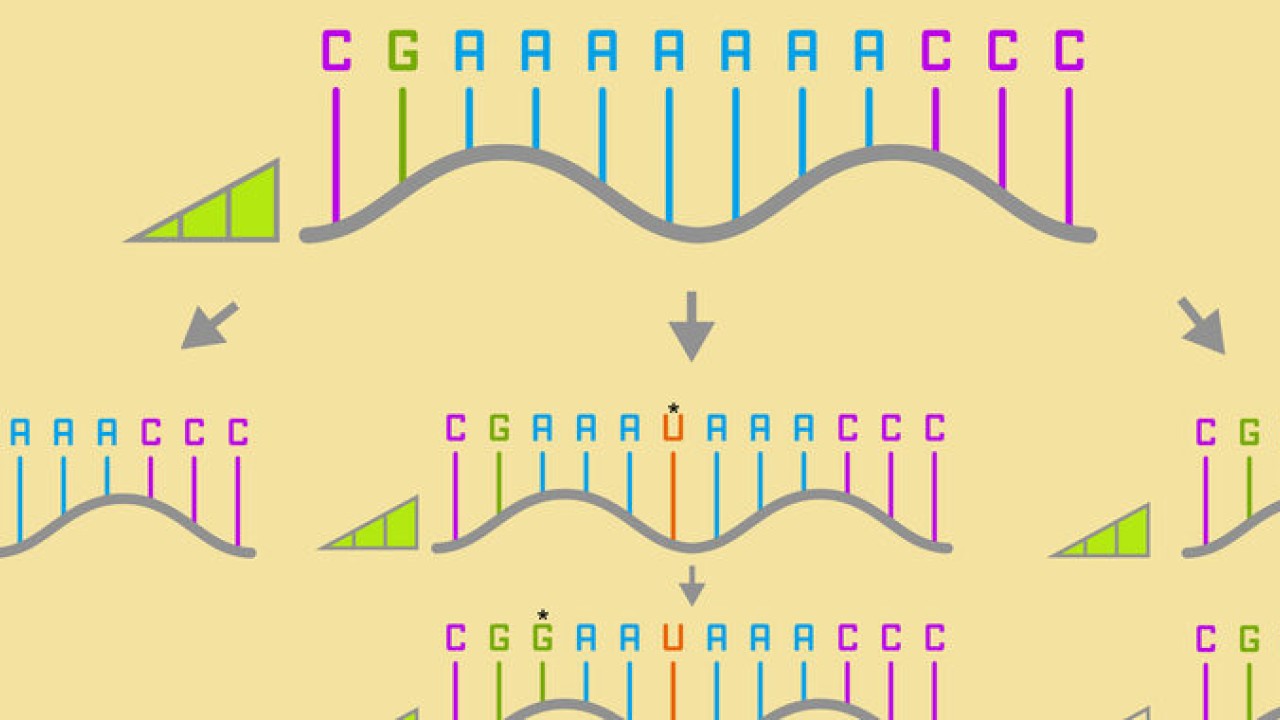
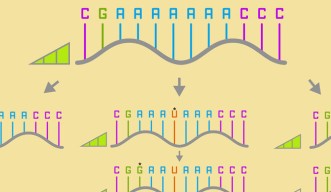
There are four standard base pairs for RNA and DNA. For RNA, these are ‘A’, ‘G’, ‘C’, and ‘U’. Prof. Yokobayashi explained the concept of a neutral network by giving an example of a simplified sequence of RNA bases.
“Say, the RNA sequence AAAAAAA mutates to AAAUAAA, which then mutates to GAAUAAA. The first variant is connected to the second one, which is connected to the third by just one single mutation. If these mutations maintain the same fitness, the organism might survive, and the mutation might be inherited by future generations. This increases the overall diversity, and diversity is essential for a species to adapt to changes in the environment.”
A neutral network is a whole series of base sequences just like this (albeit much longer), where each new sequence, differing by just one base, has roughly the same fitness as the one before and after. Scientists have suspected their existence for some time, but they’re very difficult to prove experimentally. Originally it was predicated that all RNA sequence spaces should have a possibility of harboring a large neutral network, but no one has ever found these large-scale neutral networks in practice.
In this study, the researchers looked at a type of RNA called a ligase ribozyme, which had previously been synthesized but wasn’t known to constitute a neutral network. They chose this ribozyme as its function is to connect or ligate two pieces of RNA together. This role has important implications for the origin of life as it’s vital for self-replication.
The entire ribozyme has around 80 base pairs, but the researchers focused on an area of 35 base pairs that is important for the function of the ribozyme and thus for measuring its overall fitness.
Through computational sequence design aided by an evolutionary algorithm and deep learning, the scientists designed many mutants of this ribozyme and experimentally determined their fitness. They ended up testing more than 120,000 variants and were able to increase the fraction of neutral variants in the designed population from around 10% to almost 90%.
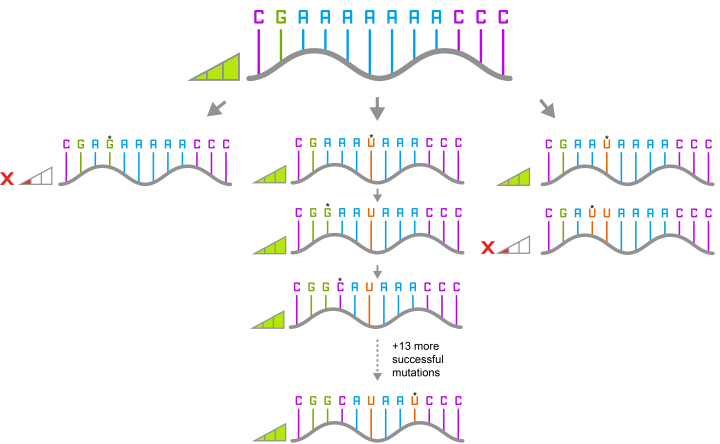
They then picked one of the variants that they had identified. This differed from the original ribozyme by 16 mutations. They screened all the different paths that could have been taken to get from the original ribozyme to the new variant, one mutation at a time for 16 steps, and found 65,536 different variants of which 60% were functional.
The researchers then looked at the paths that only contained functional mutations and found that 10% of the pathways were accessible.
“This is pretty high,” said PhD candidate and first author, Rachapun Rotrattanadumrong. “Other experimental work has never found such a large number of accessible pathways between two variants.”
The researchers emphasized that this experimental evidence of a neutral network has shown that it is possible for neutral networks to form in an RNA sequence space, though not as extensively as the theoretical work would have them believe. Another interesting question may involve looking at what properties are needed for this kind of network to form.
“This work will allow people to experimentally test many hypotheses,” continued Rotrattanadumrong. “We’ve provided the first experimental dataset of a neutral network. It will allow researchers to answer questions on how RNA evolved and continues to evolve.”
Article Information
Research Unit
Submit a press inquiry