Research Units
View by Faculty Member, Research Unit, or Research Specialties
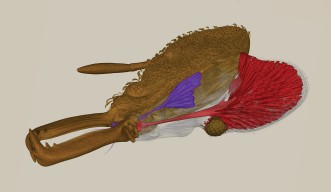
Biodiversity and Biocomplexity Unit
The Biodiversity and Biocomplexity Unit explores how ecological and evolutionary processes generate and sustain biodiversity, and how those processes are being altered by human activities.

Evan P. Economo
Professor (Adjunct)
Biological Complexity Unit
The Biological Complexity Unit studies how biophysical systems, ranging from subcellular circuits to cellular populations, can function despite being subject to random fluctuations.

Simone Pigolotti
Professor

Biological Design Unit
The Biological Design Unit investigates what determines biological forms by untangling the evolution of structural functions and organismal development in the changing climate.

Naomi Nakayama
Professor
Biological Nonlinear Dynamics Data Science Unit
The biological nonlinear dynamics data science unit investigates complex systems explicitly taking into account the role of time. We do this by instead of averaging occurrences using their statistics, we treat observations as frames of a movie and if patterns reoccur then we can use their behaviors in the past to predict their future. In most cases the systems that we study are part of complex networks of interactions and cover multiple scales. These include but are not limited to systems neuroscience, gene expression, posttranscriptional regulatory processes, to ecology, but also include societal and economic systems that have complex interdependencies. The processes that we are most interested in are those where the data has a particular geometry known as low dimensional manifolds. These are geometrical objects generated from embeddings of data that allows us to predict their future behaviors, investigate causal relationships, find if a system is becoming unstable, find early warning signs of critical transitions or catastrophes and more. Our computational approaches are based on tools that have their origin in the generalized Takens theorem, and are collectively known as empirical dynamic modeling (EDM). As a lab we are both a wet and dry lab where we design wet lab experiments that maximize the capabilities of our mathematical methods. The results from this data driven science approach then allows us to generate mechanistic hypotheses that can be again tested experimentally for empirical confirmation. This approach merges traditional hypothesis driven science and the more modern Data driven science approaches into a single virtuous cycle of discovery.

Gerald Pao
Assistant Professor

Biological Physics Theory Unit
We seek the principles governing the behavior of whole organisms, integrating physics, biology and computational approaches to understand life's most complex and fascinating phenomena.

Greg J Stephens
Associate Professor (Adjunct)
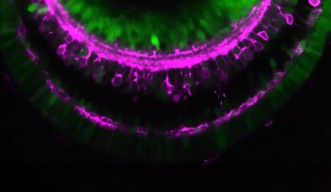
Developmental Neurobiology Unit
Developmental Neurobiology Unit uses zebrafish retina as a model to study mechanisms that control neuronal differentiation and circuit formation, and neuronal degeneration and regeneration.

Ichiro Masai
Professor

Evolutionary and Synthetic Biology
The Evolutionary and Synthetic Biology Unit focuses on understanding how living things evolved using computation, theory, experiments and field work. Combining the understanding of evolutionary mechanisms with evolutionary theory, computational and synthetic biology approaches we design novel biological objects and further penetrate the mysteries of the evolution of life.

Fyodor Kondrashov
Professor
Human Evolutionary Genomics Unit
We use the genomes of Neandertals and Denisovans, the closest evolutionary relatives of present-day humans, to identify genomic variants that are unique to modern humans.

Svante Pääbo
Professor (Adjunct)

Macroevolution Unit
Theory-driven research on macroevolution, biodiversity, and evolutionary patterns across space and time, integrating fossils, fishes, biomechanics, phylogenies, and models.

Lauren Sallan
Assistant Professor
Membranology Unit
The human body is composed of ~37 trillion cells, all of which are surrounded by a plasma membrane. We aim to understand the relationship between plasma membrane damage and multiple pathophysiological processes including aging.

Keiko Kono
Associate Professor
Microbial and Ecosystem Ecology Unit
Our unit focuses on understanding how environmental changes impact soil microbes, particularly those that are symbiotic with or pathogenic to plants, and drive biogeochemical cycling.

Chikae Tatsumi
Assistant Professor
Model-Based Evolutionary Genomics Unit
The Model-Based Evolutionary Genomics Unit works at the crossroads of computational and evolutionary biology. Our long-term goal is to achieve an integrative understanding of the evolution of Life on Earth and the origins and emergence of complexity across different biological scales, from individual proteins to ecosystems. To move towards this goal, we develop and apply model-driven evolutionary genomics methods to reconstruct the Tree of Life and the major evolutionary transitions that have occurred along its branches.

Gergely János Szöllősi
Associate Professor

Sensory and Behavioural Neuroscience Unit
Our goal is to find how the brain supports complex and adaptive behaviour, that is crucial for animals' success. We investigate this using olfaction in rodents, through study of neural circuit analyses - both local and long-range - in a variety of behavioural contexts.

Izumi Fukunaga
Associate Professor

Synapse Biology Unit
The Synapse Biology Unit studies how the dynamic features of synaptic connections between neurons mediate and maintain efficacious information processing in the brain. Synaptic communication...

Yukiko Goda
Professor





