31 January 2022
Scientists use 'sticky' DNA to build organized structures of gel blocks
Highlights
- Scientists have used strands of microscopic DNA to direct the self-assembly of hydrogel blocks that can be seen by the naked eye
- The hydrogel blocks are so far the largest objects programmed by DNA to form organized structures
- The hydrogel cubes could selectively stick to other cubes due to matching strands of DNA attached to their surface
- The researchers programmed the hydrogel blocks to self-assemble into branching structures of alternating colors and to self-sort into groups of the same color
- The team also showed that the hydrogel structures could be disassembled by disrupting the pairing between the DNA strands that were holding the blocks together
Press Release
Researchers from the Okinawa Institute of Science and Technology Graduate University (OIST) have used microscopic strands of DNA to guide the assembly of gel blocks that are visible to the naked eye.
The hydrogel blocks, which measure up to 2mm in length and contain DNA on their surface, self-assembled in around 10-15 minutes when mixed in a solution, the scientists reported today in the Journal of the American Chemical Society.
“These hydrogel blocks are, we believe, the largest objects so far that have been programmed by DNA to form organized structures,” said Dr. Vyankat Sontakke, first author of the study and a postdoctoral researcher in the OIST Nucleic Acid Chemistry and Engineering Unit.
The process of self-assembly – in which an organized structure spontaneously forms when two or more individual components interact – is common in nature, with cells and DNA able to self-assemble into amazingly complex microscopic structures. But using interactions that occur on the molecular scale to direct the assembly of macroscopic objects (meaning visible to the naked eye) is a relatively new field of research, particularly with DNA.
“We chose DNA because it is so programmable, which it owes to its exquisite ability to recognize sequences,” said senior author, Professor Yohei Yokobayashi, who leads the Nucleic Acid Chemistry and Engineering Unit.
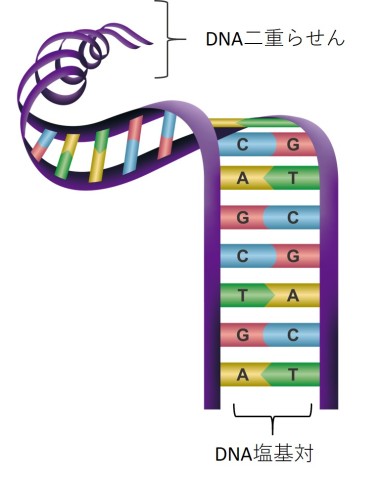
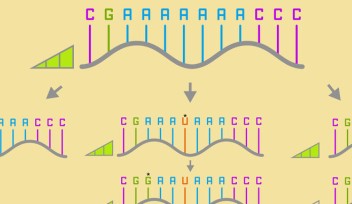
A double-stranded molecule of DNA is formed by two single strands of DNA that twist around each other to form a double helix shape. The strands are kept together by bonding between bases, which fit together like a jigsaw (A with T, and C with G). This specific base pairing ability means that scientists can design strands of DNA that match other strands precisely and will bond together.
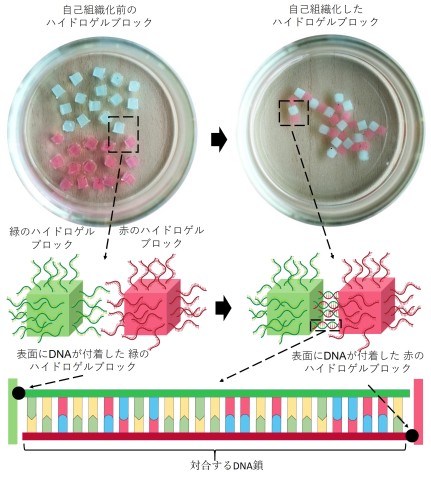
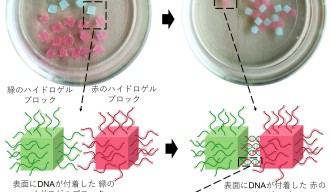
In one of the experiments, the researchers attached molecules of single-stranded DNA to the surface of red and green-colored blocks of hydrogel. The strands of DNA on the red blocks matched the strands of DNA on the green blocks.
When the hydrogel blocks were shaken in a solution, the matching strands of DNA paired together, acting like a “glue” that stuck the red and green blocks together. After ten minutes, the separated blocks self-assembled into a simple branching structure of alternating colors.
Importantly, the DNA strands did not interact with the identical strands of DNA on other blocks, so hydrogel blocks of the same color did not stick together.
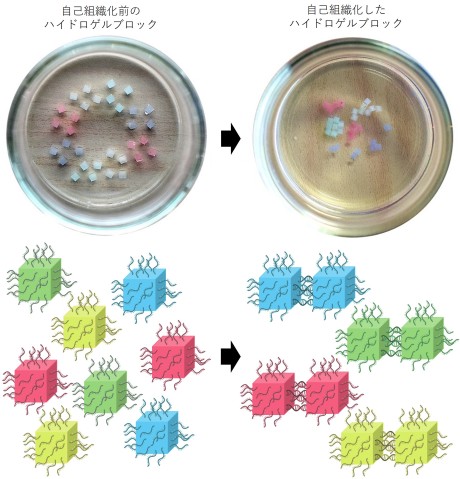
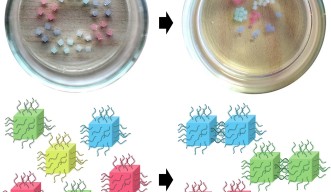
The scientists further tested the ability for the DNA to recognize only specific sequences, by designing four pairs of matching strands. They attached the single stands from the first matching pair to the surface of the red hydrogel cubes. The same process was done for the green, blue and yellow hydrogel cubes.
When shaken together, despite the presence of many different DNA sequences, the strands only bonded with their matching strand, resulting in the previously mixed up hydrogel blocks self-sorting into groups of the same color.
“This shows that the process of self-assembly is very specific and easily programmable. By simply changing the sequence of DNA, we can guide the blocks to interact with each other in different ways,” said Prof. Yokobayashi.
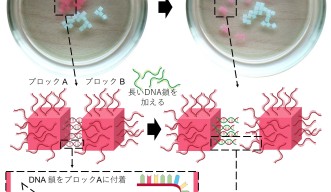
As well as self-assembly, the researchers also studied whether they could use DNA to program the disassembly of a structure. They created two matching single strands of DNA, and then made a third shorter strand that matched part of the first strand. They attached the first strand, and the matching shorter strand to hydrogel cubes, which self-assembled when mixed in solution. The longer strand of DNA that matched the first strand was then added to the solution and over the process of an hour, the longer strand displaced the shorter strand, causing the cubes to disassemble.
“This is really exciting because it means that by using DNA as the “glue” to stick the hydrogel blocks together, the process is fully reversible,” said Dr. Sontakke. “This means that the individual components can also be re-used.”
While the structures formed so far are simple, the researchers hope to add more complexity by increasing the number of different cubes that are incorporated into the structure and by targeting different DNA strands to specific cube faces. They also plan to further increase the size of the hydrogel blocks.
“This is still basic research, but in the future, these techniques could be used for tissue engineering and regenerative medicine,” said Prof. Yokobayashi. “It might be possible to place different types of cells inside hydrogel cubes, which can then assemble into the complex 3-D structures needed to grow new tissues and organs.
“But,” he added. “Regardless of potential applications, it’s incredible to be able to witness chemistry as microscopic as interacting DNA strands with our own eyes. It’s a really fun piece of science.”
Research Article
Title: Programmable Macroscopic Self-Assembly of DNA-Decorated Hydrogels
Journal: Journal of the American Chemical Society.
Authors: Vyankat A. Sontakke and Yohei Yokobayashi*
Date: 31st January 2022
DOI: doi.org/10.1021/jacs.1c10308
Research Unit
Submit a press inquiry