Dr. Yoshiyuki Asai Awarded for Work on PhysioDesigner Software
In the early days of biological research, organisms were examined from a purely behavioral perspective; it was not until molecular biology made an appearance that scientists were able to pin down some of the mechanisms behind cellular processes. But even almost a decade after the last base pairs of the human genome were identified, biologists still lack a clear, overall understanding of how all the components of organisms fit together.
To many, it seems that the solution lies in an area of science that OIST's Prof. Hiroaki Kitano was amongst the first to describe. The emerging field of systems biology takes a holistic approach, aggregating knowledge from diverse components to form a complete model of an organism. By defining sets of differential equations, it is possible to reproduce biological phenomena such as the diffusion of ions across a cell membrane. Combining these mathematical models then allows scientists to 'build up' a model of an organism.
But combining multiple models like this is tricky - especially for a complex organism such as the human body, which requires a hierarchy of models to describe phenomena at the levels of genes, proteins, cells and organs.
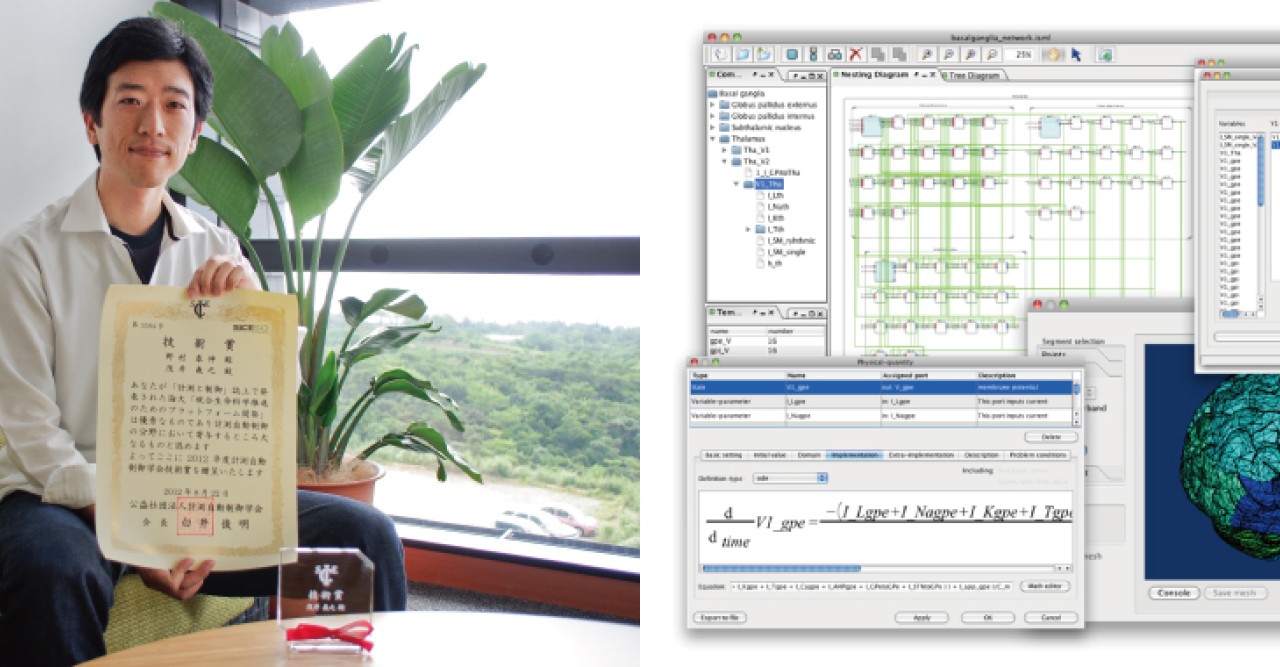
This is where Dr. Yoshiyuki Asai of Prof. Kitano’s Open Biology Unit comes in. Along with other members of his unit, he has developed a new software platform, PhysioDesigner, to assist those who want to create physiological models. It is for his work on the program that Dr. Asai was presented with an award by the Society of Instrument and Control Engineers (SICE) in late August.
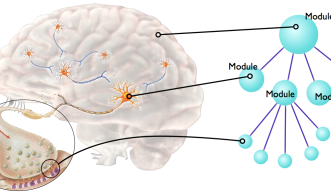
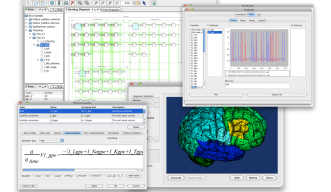
PhysioDesigner was called insilicoIDE when Dr. Asai joined the project as a chief developer in 2007 at Osaka University. After Dr. Asai’s joining OIST, Dr. Kitano and he rebranded it with major reengineering, and released it as PhysioDesigner. It enables researchers to construct biological systems from hundreds or thousands of 'modules', and to describe the interactions between them with mathematical equations. Each module can then be sub-divided into another series of modules and so on, thus creating the required hierarchical structure.
The software stores the models in Physiological Hierarchy Markup Language (PHML), an XML-based format, but by also supporting other languages such as Systems Biology Markup Language (SBML), it allows researchers to share and merge models. Once a model has been constructed, it is then passed through PhysioDesigner’s sister simulation program, Flint, which generates the required numerical data.
The potential for modeling physiological systems in this way is enormous, particularly for medicine and pharmaceutical research. "Currently the medical treatments, such as dispensing of drugs, are decided based on medical trials and the experiences of doctors," explains Dr. Asai. "If we can construct appropriate models, we can calculate how the chemical compounds in the drug will interact to the proteins in the cell, and then predict the overall effect on a patient. PhysioDesigner can contribute to promote such activities."
PhysioDesigner is available at physiodesigner.org. Please try it. Any feedback is appreciated.
Specialty
Research Unit
For press enquiries, please contact media@oist.jp